Introduction
ChallengeSimple Western™ Jess is a high throughput western blot technique with truly quantifiable data. However, preparing the samples and Simple Western™ plate is time consuming, laborious, and is prone to small errors.
SolutionThe CyBio FeliX was used to perform all liquid handling aspects of the sample protein assay and the Simple Western™ plate preparation.
The Simple-Western™ Jess is a capillary based automated western blot instrument capable of running 24 samples from generation to results in approximately 5 hours. This throughput is advantageous compared to traditional western blot systems that traditionally runs 10-20 samples, has 4-5 hours bench time, and takes 24-48 hours to get results.
Automation of the protein separation and immunodetection process eliminates the error prone steps of traditional western blotting and the associated software quickly processes the quantifiable data. At Aurelia Bioscience, the Simple-Western™ Jess is a staple technique in a repertoire of drug discovery methodologies.
For this reason, the CyBio FeliX liquid handling system was used to simplify the Simple-Western™ set-up procedure.
Automation of the liquid handling aspect of Jess plate preparation would further increase the reproducibility of the data, allow increased flexibility of a scientist’s workload by removing the manual aspects of the preparation, and allow inexperienced users to access the technology. To demonstrate the capabilities of CyBio FeliX, THAL-SNS-032, a well-studied Proteolysis-targeting chimera (PROTAC) molecule and specific degrader of cyclin dependent kinase 9 (CDK9) was used to demonstrate the capability of the system and compare the results to previously generated data used for the Protein-Simple™ webinar “Advancing Targeted Protein Degradation Research with Automated Western Blotting”.
Materials and Methods
Samples and reagents
- Pierce BCA protein assay kit – Thermo Fisher #23227 (Reagents B and A are mixed at a 1:50 ratio prior to use)
- BCA reagent A, 500 ml
- BCA reagent B, 25 ml
- 12-230kDa Jess/Wes separation module 8x25 capillary cartridges – Bio-Techne #SM-W004
- Anti-Rabbit Detection module for Jess/Wes – Bio-Techne #DM-001
- Anti-Mouse Secondary NIR antibody – Bio-Techne #043-821
- MV-4-11 Cells – ATCC #CRL-9591
- RPMI-1640 – Thermo Fisher #21875-034
- Fetal Bovine Serum – Corning #35-079-CV
- PBS tablets – Thermo Fisher 18912014 (1 tablet per 500 mL sterile water)
- 1x Complete RIPA lysis buffer:
- RIPA – Thermo Fisher #89900
- Magnesium Chloride – Merck #M1028-100 ml (FAC 2mM)
- Benzonase clease – Merck #E1014-25KU (0.5 µL per mL)
- Pierce Protease Inhibitor Tablets, EDTA-free – Thermo Fisher #A32965 (1 tablet per 50 mL lysis buffer)
- THAL-SNS-032 – Tocris #6532
- Anti-CDK9 antibody – Cell Signaling Technology #2316
- Anti-β-actin antibody – Bio-Techne #MAB8929
Instrumentation and Consumables
- CyBio FeliX Basic Unit with Enclosure (Analytik Jena GmbH) (OL5015-24-100)
- CyBio FeliX CHOICE Head (OL3316-14-250)
- 8-channel CHOICE Adapter; 10-1000 µL (OL3316-14-330)
- 8-channel CHOICE Adapter; 1-50 µL (OL3316-14-332)
- 2 x Adapter 24 tubes, passive cooling function (844-00136-0)
- CyBio Waste box (844-00430-0)
- CyBio TipBox 96/250 µL Sterile (OL3811-25-637-S)
- CyBio TipBox 96/50 µL Non-sterile (OL3811-25-535-N)
- Adapter 96, passive cooling function (844-00135-0)
- 2 x Height adapter 40 mm (844-00445-0)
- 12-channel Reservoir Plate – Alphalabs #LW9400
- Simple-Western Jess
- Perkin-Elmer EnSpire – Reading plate at 562 nm
- 1.5 mL Eppendorf Tubes – SLS #E0030123328
- PCR tube strip and lids – Alphalabs #LW2535
- 24-well plate – Thermo Fisher #930186
- Polystyrene clear 384-well plate – Greiner #781101
- Plate Seals – Alphalabs #LW2770
- Bio-Rad MyCycler thermal cycler
- Thermo Scientific Megafuge 8R + M10 microplate swinging bucket rotor
- 37 °C incubator – Stuart Scientific incubator SI 18
Cell treatment and lysate preparationMV-4-11 cells are seeded in a 24-well sterile polystyrene plate and incubated at 37 °C with 5% CO2 and humidity.
The cells are treated with serially diluted SNS-THAL-032 in technical duplicate before being collected in 1.5 mL tubes. The cells are washed in PBS before being lysed in 1x complete RIPA lysis buffer, incubated on ice for 30 minutes then clarified at 17,000 × g for 10 minutes at 4ºC.
Bicinchoninic Acid (BCA) Assay (CyBio FeliX)
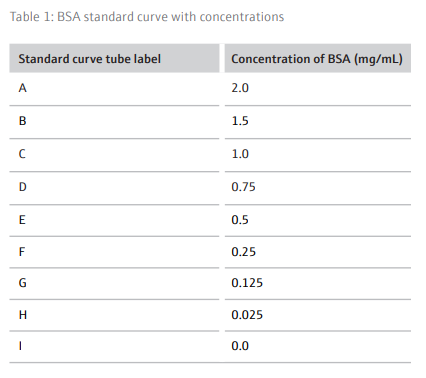
The BCA assay is a method of determining a cell lysate’s protein concentration by extrapolating unknown values from a standard curve constructed from samples with a known concentration of a specific protein (in this case BSA) in an appropriate buffer. For this assay BSA concentrations
ranging from 2.0-0.025 mg/mL (Table 1) were used to determine the unknown protein concentrations from 24 unique samples at a time.
Prior to running the BCA assay, the CyBio FeliX prepared the dilution series of protein standards for use across multiple BCA assays in a separate protocol. The CyBio FeliX performs the assay by following plate template and the instructions below:
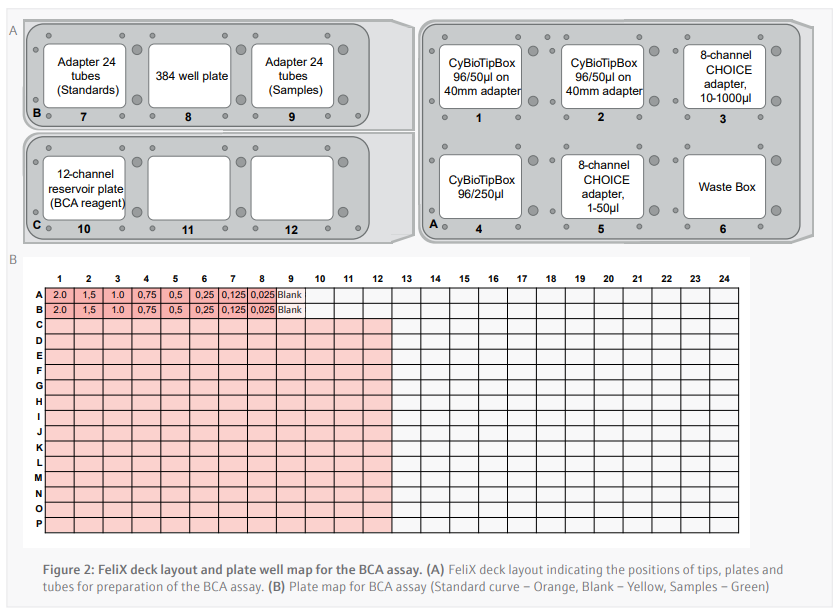
- 16 µL of each protein standard is added in duplicate across wells A1-A9 and B1-B9 of a polystyrene 384-well plate (Figure 2B).
- A 1/3 protein sample dilution is carried out in the well using a single channel and 50 µL tips as described below:
- 4 μL of lysis buffer is added to a designated well of the 384-well plate in duplicate.
- b. New tip is picked up and 2 μL of sample is then be added to each of the duplicate wells (final volume of 6 μL per well).
- 50 µL of pre-mixed BCA reagent is added to each well
Upon completion the following steps were performed outside the FeliX:
- Plate is sealed by hand.
- Incubation of the plate at 37 °C for 30 minutes.
- Read absorbance at 562 nM on the EnSpire plate reader.
To construct the standard curve, the standards of known BSA concentration are plotted against their respective absorbances. Linear regression is then used to calculate the protein concentrations of cell lysate samples with an unknown protein content from their absorbances. An average of the duplicate samples is taken then multiplied by the dilution factor used.
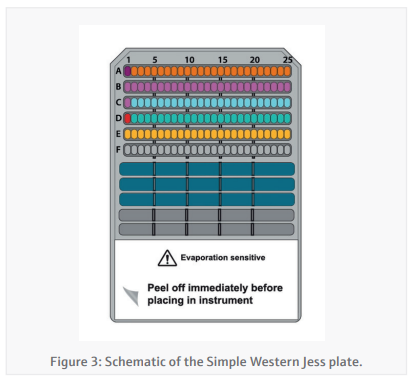
Simple-Western Jess plate preparation (CyBio FeliX)By providing the CyBio FeliX with the Jess plate template generated in the Simple-Western Compass software and the protein concentrations generated in the BCA assay, the preparation of the Jess plate (Figure 3) is almost entirely autonomous. The CyBio FeliX performs the plate set-up as
described in the instructions below:
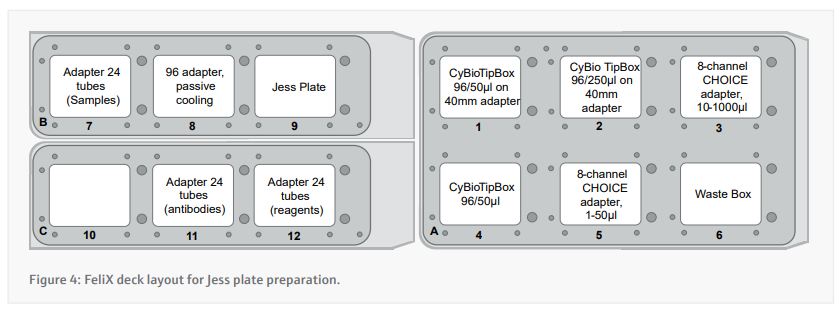
- Prepare the deck appropriately according to the instructions presented by the CyBio FeliX. A layout of where to place items on the deck is included in figure 4.
- Each cell lysate is diluted to the desired concentration using a variable volume of 0.1x Sample buffer and a fixed volume of cell lysate in PCR tube strips.
- Small tubes of DTT and master mix are reconstituted using the appropriate volumes before dispensing 1.25 µL into fresh PCR tube strips.
- Transfer 5 µL of diluted cell lysate into the 1.25 µL of master mix.
- These PCR tube strips now need to be taken off FeliX to a Thermocycler to be kept at 95 °C for 5 minutes using the thermocycler. The next steps up to 9 can continue while this takes place.
- The small tube of biotinylated ladder is reconstituted and 10 µL transferred into well A1.
- 10 µL of Streptavidin-conjugate is transferred into well D1.
- The following steps are carried out by multi-dispense using the 8 Channel CHOICE 10-1000µl adapter and a single 250 µL tip (tips swapped for each reagent)
- 10 µL of Antibody dilution buffer is transferred into wells B1-B25 and C1.
- 10 µL of Primary antibody solution is transferred into the appropriate wells between C2 and C25.
- 10 µL of Secondary antibody solution is transferred into the appropriate wells between D2 and D25.
- If appropriate, 15 µL of pre-mixed 1:1 Peroxide : Substrate is added to wells E1-E25.
- The PCR strips are returned to the cool block in the FeliX and 4 µL of the samples is transferred into the corresponding wells between A2 and A25
- 500 µL of wash buffer is added into all wells of row G, H, and I.
Before running the plate on the Simple-Western Jess, the plate in centrifuged at 1250 × g for 5 minutes and alcohol vapour is applied using a squeeze bottle with the internal straw removed to ensure removal of any bubbles. At this point the plate is run on the Jess instrument using the template presented to the CyBio FeliX at the initial phase of set-up.
Results and Discussion
Antibody optimization (CyBio FeliX) Due to the sensitivity and quantitative nature of the Simple-Western Jess, each antibody must not only be optimized for its target, but this optimization is also specific to each cell line. Antibody saturation and concentration dependent signal linearity are the two aspects of antibody optimization that must be achieved to ensure the data received is truly quantifiable.
To determine the activity of THAL-SNS-032, the anti-CDK9 antibody and the anti-β-Actin was optimized by using the CyBio FeliX. Using an MV-4-11 lysate of pre-determined protein concentration, the CyBio FeliX used serially diluted lysate and varying antibody dilutions to prepare the Simple[1]Western Jess plate, completely removing the hands-on steps normally required for optimizing an antibody.
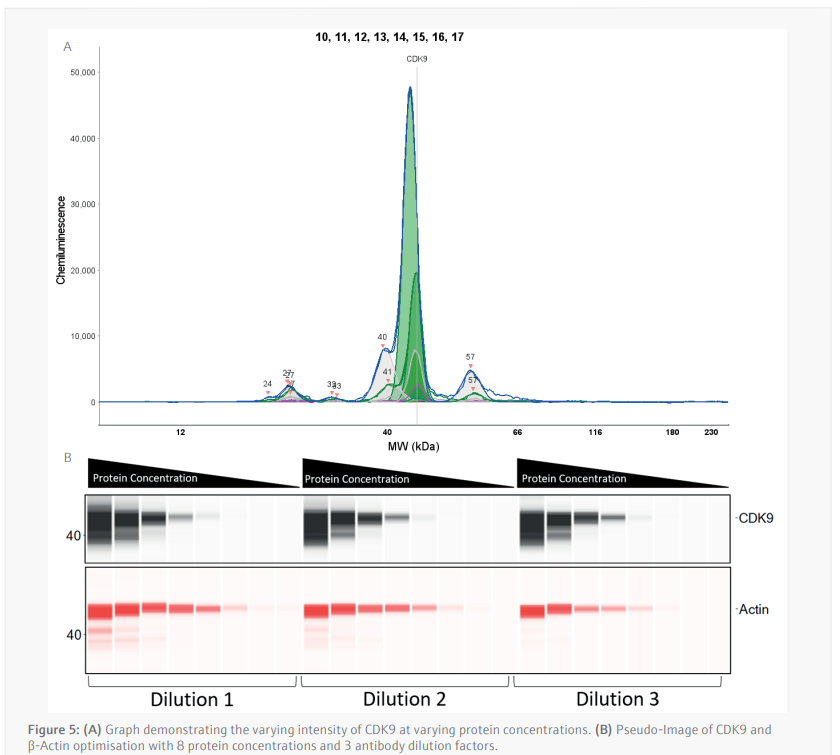
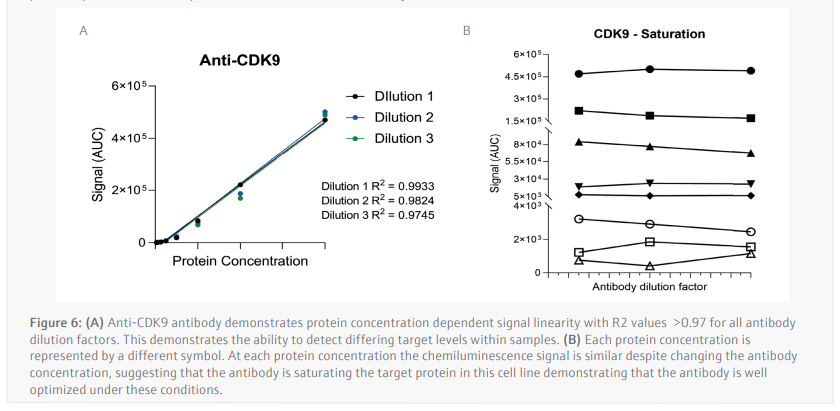
The luminescence or near-infrared (NIR) signal emitted from the glass capillary by the secondary antibodies is detected by the camera within the Jess and is transformed into an intensity graph (Figure 5A). A pseudo-image representation of the graphs can be created to give a traditional style visualization of the data (Figure 5B). The peak height or area under the curve (AUC) can then be used to determine the linearity and saturation of the antibody. A graph of the AUC of CDK9 signal on the Y-axis against protein lysate concentration on the X-axis can then be plotted, the R2 of the line of best fit is a representation of the linearity of the antibody (Figure 6A). The linearity allows to determine if the antibody can detect changes in target protein within the sample. A graph plotting the AUC of CDK9 on the Y-axis against antibody dilution factor on the X-axis for each protein concentration visualizes the ability of the anti[1]CDK9 antibody to saturate its target (Figure 6B). Antibody saturation evaluates whether all the epitopes of the target (in this case CDK9) are being occupied despite changes in antibody concentration. An example of sub-optimal saturation would be if changes in signal are seen across the different antibody dilution factors, this indicates that all the target epitopes are not being occupied. The optimization of the CDK9 antibody demonstrates the signal remains the same across all antibody concentrations, indicating that antibody concentration will not be a factor in the CDK9 signal present during any subsequent experiments. The data shown suggests that the anti-CDK9 antibody is linear with an R2 >0.97 and saturated across all antibody dilution factors and protein concentrations (no antibody concentration dependent changes).
This analysis was also carried out for the anti-β-Actin antibody in the NIR channel which yielded similar results. This allowed the selection of a specific protein concentration and antibody dilution factor for each antibody.
Jess plate preparation (CyBio FeliX)
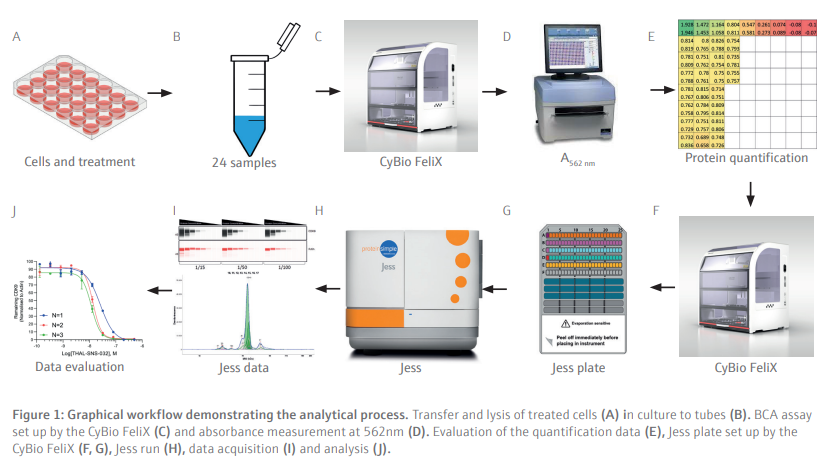
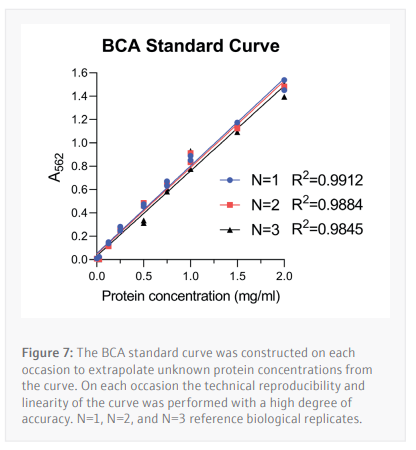
Following optimization of the antibody dilution factors and cell line protein concentrations, cells are seeded then treated with serially diluted THAL-SNS-032 (500 nM top concentration with a 2/5 dilution factor) before being washed with PBS then lysed with RIPA lysis buffer. The CyBio FeliX is then used to perform the BCA protein assay, increasing reliability, and removing the hands-on requirements of assaying protein concentration for each of the 24 lysates generated (Figure 7). This step is important to ensure that the cell lysates can be diluted to the appropriate concentration identified during the antibody optimization steps carried out by the CyBio FeliX previously. Once the unknown protein concentration of samples has been determined the Simple-Western Jess plate can be prepared by the CyBio FeliX in an accurate and timely manner.
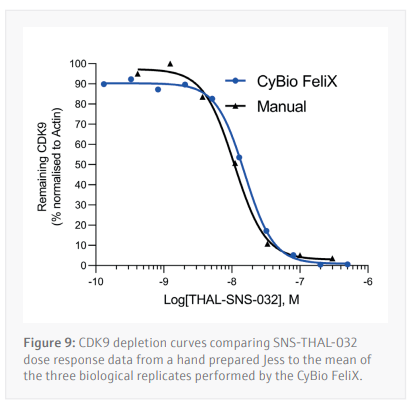
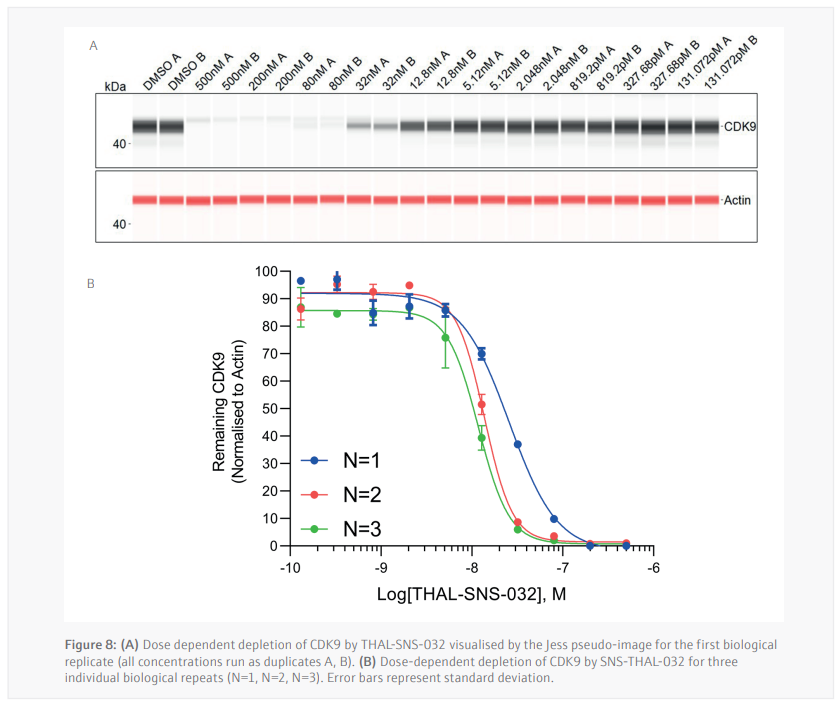
Three biological replicates were performed to determine the reproducibility for the Jess plate preparation. The peak’s intensity data generated from each capillary is used to visualize the data in a pseudo-image format (Figure 8A). Quantification of these peaks can then be performed and plotted as a dose response curve (Figure 8B). In all three biological replicates SNS-THAL-032 demonstrated a potent and reproducible dose dependent degradation of CDK9 with a DC50 of 16.1 nM ± 7.6 nM. Good technical reproducibility for each concentration is also demonstrated for all three biological replicates, giving further evidence of the accuracy and reliability of the CyBio FeliX. Furthermore, comparison of Simple Western Jess plate preparation by the CyBio FeliX to a historical, manually prepared Jess, demonstrated the ability to replicate data while saving a scientist up to 60 minutes of bench time per Jess run (Figure 9).
 Conclusion
Conclusion The Simple-Western™ Jess expertise at Aurelia Bioscience combined with the liquid handling automation expertise at Analytik Jena lead to the development of a fully automated western blotting methodology. The CyBio FeliX processes cell lysates from the protein assay through to quantifiable results, reducing bench-time by 60 minutes per Jess run.
Using the PROTAC THAL-SNS-032, a potent degrader of CDK9, we demonstrated that the CyBio FeliX is a flexible and reliable liquid handling platform which is able to assist with the preparation of the Simple Western Jess plate and quantify changes in CDK9 levels within a sample. With these PROTAC assays it is absolutely critical that we can reliably and quantifiably determine a targets level within a given sample, this alone places a large emphasis on liquid handling accuracy, something that the CyBio FeliX is well suited for. With this methodology we can now apply the use of this platform to optimize the Simple Western Jess assay for new targets through reliably optimizing antibody and sample protein concentrations. This hands-off approach adds flexibility to a scientist’s workload and allows less experienced users to access the technology.